|
|
|
MicrobeTracker
|
 |
|
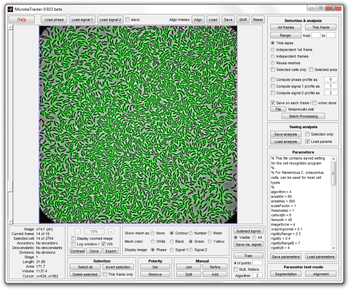
Main window of MicrobeTracker |
MicrobeTracker is a MATLAB-based program designed to detect and outline
bacterial cells in images obtained by light microscopy and to analyze
fluorescence signal inside them. The program generally requires phase contrast
images for the analysis, though in some cases they may be replaced with
fluorescence images, e.g. with diffuse fluorescent proteins, or with DIC images
that have to be converted to symmetric (phase contrast-like) representation
using a tool included in a separate file. When applied to individual images,
MicrobeTracker combines traditional thresholding technique with morphological
opening operations and edge detection to obtain the initial guess of the shape.
Then this shape is additionally Fourier-smoothed and may be refined using one of
a few more complicated techniques, including versions of Active Contour and
Point Distribution Model (PDM).
Detected cells are saved in the form of 'meshes', which
provide a coordinate system along the cell. The meshes alone can be used to
measure the cell diameter and its uniformity, and to determine cell length, volume,
curvature and other geometrical parameters. In combination with fluorescence
images, the program can be used to determine the profile of the signal along the
cell. This signal provides the total intensity and relative intensity of individual
fluorescence foci or polarity of localization.
The program is written in MATLAB and C/C++, it requires
MATLAB to run and saves the data in MATLAB (.mat) format. It also requires
MATLAB's Image Processing toolbox.
Contents
|
|
|