|
The panel shown above is located in the lover right corner of
MicrobeTracker window. The mode is designed to display normally hidden
intermediate images for the purposes of choosing the optimal parameters for a
particular image. There are two parameter testing regimes: Segmentation and
Alignment. Segmentation mode will preview the segmented image before the
alignment started with the current parameter set (the only mode is algorithm 1
is selected), while Alignment mode will display the process of the active
contour fitting to each cell.
Segmentation
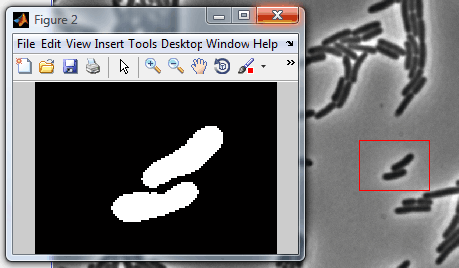 This
mode previews the segmented image for the current frame before the contour
alignment starts and opens a separate MATLAB figure with the result. If
Selected area is chosen, it only processes this area (see the image to the
right). If no area is selected, the whole image gets processed. This
mode previews the segmented image for the current frame before the contour
alignment starts and opens a separate MATLAB figure with the result. If
Selected area is chosen, it only processes this area (see the image to the
right). If no area is selected, the whole image gets processed.
This regime consists of three steps, each of the affected only by a small number
of parameters.
1. Thresholding:
2. Edge/valley detection:
-
edgemode
─ "none" (or 0) - no edge detection, "log" (or 1) - Laplacian of Gaussian (LoG),
"valley" (or 2) - Valley detection, "logvalley" (or 3) - both;
-
edgeSigmaL
(parameter for LoG algorithm)
-
edgeSigmaV
(parameter for Valley detection)
-
valleythresh1
(parameter for Valley detection)
-
valleythresh2
(parameter for Valley detection)
-
thresFactorM
(thresholding parameter)
-
edgedetection, edgeSigma, logmode ─ obsolete parameters.
3. Splitting regions:
-
splitregions
(parameter that determines whether a large region needs to be split)
-
areaMin
(smallerst area of a region in sq. pixels, all smaller regions are discarder)
-
areaMax
(largest area of the region in sq. pixels, all larger regions are split if
splitregions is selected or discarder otherwise)
In algorithm 1, every segment of
the correct size range is outlined directly. In the rest of the algorithms, the
segmentation is followed by contour creation and alignment.
Alignment
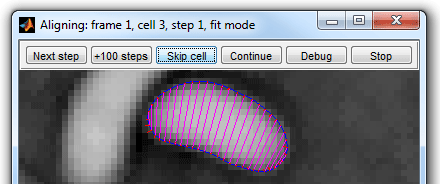 The Alignment button
on the Parameter test mode panel lets the user to see the process of alignment dynamically.
After activating the mode, click any processing button (such as All frames,
This
frame, Range, buttons for manual operations) to see the how alignment happens.
Note, MicrobeTracker still saves the data in memory and to the disk after each
frame (if selected), so be careful to not erase your data! The Alignment button
on the Parameter test mode panel lets the user to see the process of alignment dynamically.
After activating the mode, click any processing button (such as All frames,
This
frame, Range, buttons for manual operations) to see the how alignment happens.
Note, MicrobeTracker still saves the data in memory and to the disk after each
frame (if selected), so be careful to not erase your data!
This regime when activated is equivalent to
fitDisplay and
fitDisplay1
parameters present and set to 1. While fitDisplay
allows the user to see the alignment process to the image,
fitDisplay1 allows for seeing alignment to the
roughly segmented region in algorithms 2 and 3 only (it will have no effect in
algorithm 4). The alignment procedure is different depending on what
cell outlining algorithm is used. For example,
since there is no alignment in algorithm 1, these parameters and the Alignment
button will have no effect.
Buttons of the alignment tool:
-
Next step - proceed one step and display the new outline
-
+100 steps - perform 99 steps silently and display the
outline of the 100's one
-
Skip cell - finish the alignment of this cell silently
-
Continue - finish the alignment of this cell displaying
every step without the need to press any button again
-
Debug - stop microbeTracker in debug mode (for example, to
see the actual values of the forces)
-
Stop - terminate the process and exit MicrobeTracker
(equivalent to Ctrl+C keyboard combination)
The
color codes used to display the outline and forces for algorithm 4 are shown on the second
figure. Algorithms 2 and 3 use only the image forces, the constrains are
implemented in a different way and are not displayed.
|